4.3 Statistical analysis of the trajectory data by Python
Contents
Note: this tutorial is a simple introduction to using python (version 3.x) and packages such as NumPy and matplotlib to do some fundamental statistical analysis of simulation results. If you want to use python to parse MD trajectories, please try specific libraries such as MDTraj and MDAnalysis, which are beyond the scope of the current tutorial. Please refer to the website of python for a basic programming guide and get help from PyPI or anaconda about how to install packages.
Preparation
Please download the files for tutorial-4.3.
# Download the tutorial file
$ cd ~/GENESIS_Tutorials-2022/Works
$ mv ~/Downloads/tutorial22-4.3.zip ./
$ unzip tutorial22-4.3.zip
# Let's clean up the directory
$ mv tutorial22-4.1.zip ./TRASH
# Let's take a note
$ echo "tutorial-4.3: Trajectory analysis using python" >> README
$ cd ./tutorial-4.3
$ ls
Analysis1 Analysis2.1 Analysis2.2
1. Basic statistics
We first learn to do some basic statistical analysis, such as calculating the average and standard deviation and finding out the maximum/minimum values. These tasks can be easily accomplished by writing python scripts. In the following examples, we will analyze the energy data obtained in Tutorial 3.1.
# Change directory for statistics analysis
$ cd Analysis1
$ ls
energy.log 01_list.py 02_numpy.py
1.1 Using the “list” data structure
In this part, we show how to use the basic list structure to store data read from a file and how to compute the average and standard deviation.
We use the script 01_list.py to do everything. You can take a look at the file by running cat 01_list.py:
#!/usr/bin/env python3
# analyze the 5th column
i_col = 5
# use the list ene_data to store energy values
ene_data = []
with open("./energy.log", "r") as ene_file:
for line in ene_file:
ene_value = float(line.split()[i_col - 1])
ene_data.append(ene_value)
# print length of the list
len_ene_data = len(ene_data)
print("Length of the energy list:", len_ene_data)
# find out the max/min
print("Maximum of column {0:>2d}: {1:>12.4f}".format(i_col, max(ene_data)))
print("Minimum of column {0:>2d}: {1:>12.4f}".format(i_col, min(ene_data)))
# calculate the average
ene_ave = sum(ene_data) / len_ene_data
print("Average of column {0:>2d}: {1:>12.4f}".format(i_col, ene_ave))
# calculate the standard deviation
ene_std = (sum((e - ene_ave)**2 for e in ene_data) / len_ene_data)**0.5
print("Standard deviation of column {0:>2d}: {1:>12.4f}".format(i_col, ene_std))
i_colis an integer variable to specify the column of interest.ene_datais the list we use to store energy data.- all the data is read from
"./energy.log"within theforloop:lineis a “string” variable to store every line in the file;line.split()simply splits the line into a list of words;float()is a function to convert a “string” into a “float”;append()is a function to add new values to the end of theene_datalist.
len()is a function used to get the length of a list.max()andmin()are the functions used to find out the maximum and minimum values from the list, respectively.sum()is a function used to calculate the sum of a list.- in the calculation of the standard deviation, we use a python tactic called “list comprehension” to quickly construct a new list:
(e - ene_ave)**2 for e in ene_data, whose elements are the squared deviations from the average. format()inserts the values saved in a variable to the message to be printed out.
Now let’s execute the script:
$ ./01_list.py # or: python3 01_list.py
Length of the energy list: 1000
Maximum of column 5: 7.3745
Minimum of column 5: -10.2807
Average of column 5: -1.5140
Standard deviation of column 5: 2.8306
1.2 Using the “numpy.ndarray” data structure
In this part, we show an alternative way to get the maximum/minimum, average, and standard deviation values using the numpy library. numpy is a Python “package” or “module” that provides a multidimensional array object and various functions for fast operations on arrays.
The program is in the 02_numpy.py file:
#!/usr/bin/env python3
import numpy as np
# analyze the 5th column
i_col = 5
# use array ene_data to store energy values
ene_data = np.loadtxt("./energy.log", usecols=i_col - 1)
# print length of the list
len_ene_data = ene_data.size
print("Length of the energy list:", len_ene_data)
# find out the max/min
print("Maximum of column {0:>2d}: {1:>12.4f}".format(i_col, ene_data.max()))
print("Minimum of column {0:>2d}: {1:>12.4f}".format(i_col, ene_data.min()))
# calculate the average
ene_ave = ene_data.mean()
print("Average of column {0:>2d}: {1:>12.4f}".format(i_col, ene_ave))
# calculate the standard deviation
ene_std = ene_data.std()
print("Standard deviation of column {0:>2d}: {1:>12.4f}".format(i_col, ene_std))
Compared with 01_list.py, this new script looks much simpler. Actually, in many cases, numpy can simplify the development of programs. We list out the main differences between the two versions in the following:
import numpy as npis used to declare the usage ofnumpy.np.loadtxt()is a function used to read formatted data from file; the return value is an array (“ene_data“).sizeof an array is the number of elements (be aware thatsizeis not a function).max()andmin()are now used as member functions of the arrayene_data.mean()calculates the average of an array.std()directly computes the standard deviation.
Let’s run this new script:
$ ./02_numpy.py # or: python3 02_numpy.py
Length of the energy list: 1000
Maximum of column 5: 7.3745
Minimum of column 5: -10.2807
Average of column 5: -1.5140
Standard deviation of column 5: 2.8306
As can be seen, the results are the same as the ones we computed with the 01_list.py.
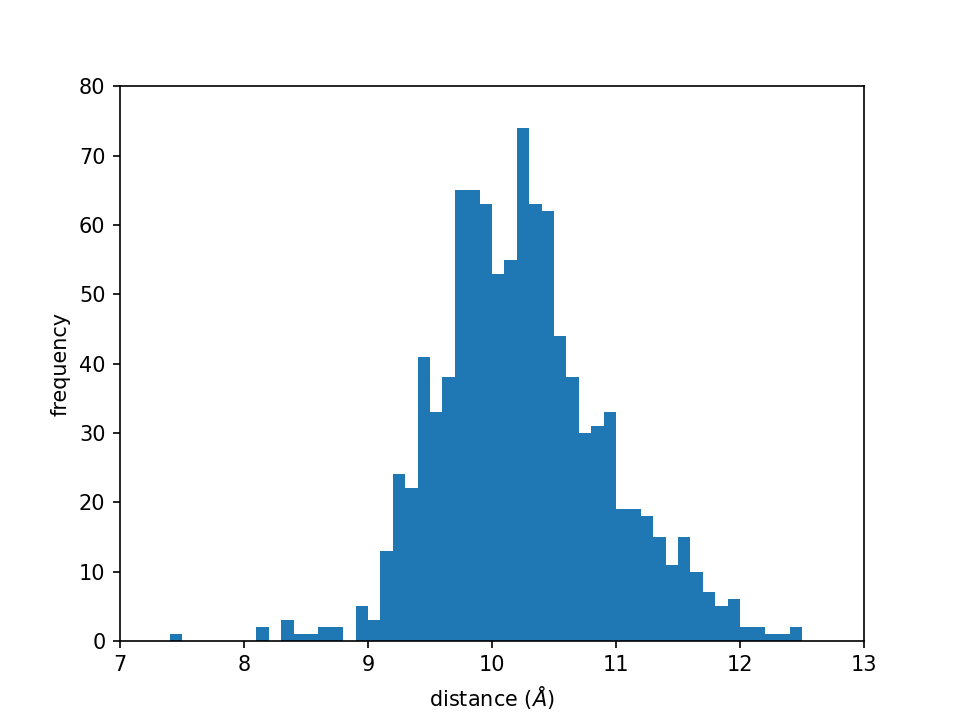
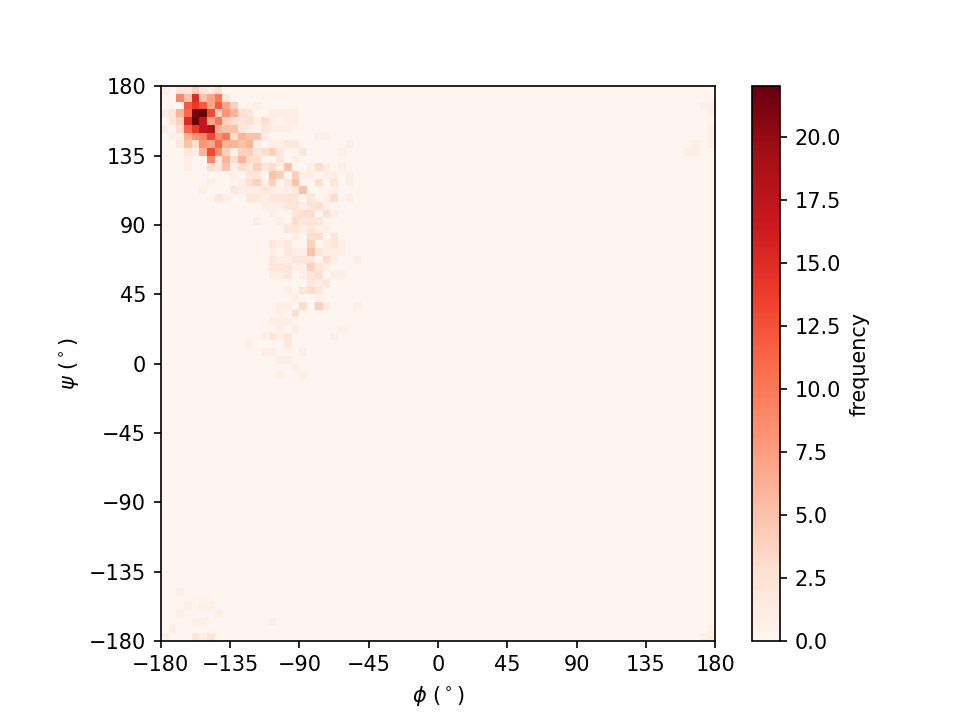
2. Histogram analysis
A histogram is an approximate representation of the distribution of numerical data. In section 2, we are going to compute and plot one-dimensional (1D) and two-dimensional (2D) histograms from the MD simulation results.