18.1 Coarse-grained MD simulation of protein with the AICG2+ model
Contents
(This tutorial is for GENESIS v1.7.0 and later)
The atomic interaction-based CG model, version 2+ (AICG2+) [1] is a variation of the classic Go-model [2] for proteins. The model represents every amino-acid by one coarse-grained particle.
Here we show an example of simulating a small protein with this model in GENESIS. We will go through all the steps from setting up the system to doing some easy analysis of the simulation results.
0. Preparations
0.1. Download GENESIS-CG-tool for preparing CG simulation files
First of all, let’s get started with a collection of scripts that helps us prepare the topology and coordinate files for GENESIS CG MD simulations. Hereafter we will call this tool as the “GENESIS-CG-tool”. It is included as a part of GENESIS, located in the subdirectory src/analysis/cg_tools. You can also find it from github and download it with git:
git clone https://github.com/noinil/genesis_cg_tool.git
This tool is developed in the programming language Julia. Therefore, you have to first install and learn some basic operations of Julia. Please also install the Julia package “ArgParse”, which is a dependency of the GENESIS-CG-tool:
$ julia
julia> import Pkg
julia> Pkg.add("ArgParse")
You can also run the following command to install dependencies:
/home/user/genesis_cg_tool/opt/dependency.jl
Remember that you can always find the help information of the GENESIS-CG-tool by running:
/home/user/genesis_cg_tool/src/aa_2_cg.jl -h
0.2. Download files for this tutorial
Now, let’s download the tutorial file (tutorial19-18.1.zip). This tutorial has three steps: 1) system setup, 2) MD simulations, and 3) trajectory analysis.
# Download and unarchive the tutorial files
$ cd /home/user/GENESIS/Tutorials
$ unzip tutorial19-18.1.zip
$ cd tutorial-18.1
$ ls
01_setup 02_simulation 03_analysis
1. Setup
We get started from preparing the topology and coordinate files for the system. Unlike atomistic simulations, we cannot simply download the PDB file and make use of some standard force-field files. Instead, we have to generate all the files by ourselves, which can be easily done with the help of the GENESIS-CG-tool.
As an example, we download a pdb file of protein GB1 from the RCSB:
$ cd 01_setup
$ wget https://files.rcsb.org/download/1PGB.pdb
We then use the GENESIS-CG-tool to extract information from the pdb file and generate the CG files:
$ /home/user/genesis_cg_tool/src/aa_2_cg.jl 1PGB.pdb
$ ls
1PGB.pdb 1PGB_cg.gro 1PGB_cg.itp 1PGB_cg.top
Don’t forget to change “/home/user/genesis_cg_tool” in the command above to your real path of the GENESIS-CG-tool.
The “1PGB_cg.gro” file stores the coordinates of the CG particles. The “1PGB_cg.itp” file contains the parameters for all the bonds, angles, dihedral angles, and the native contacts. The “1PGB_cg.top” file is the main topology file and integrates (see the “#include” lines in the file) parameters in the “1PGB_cg.itp” and other standard parameter files. Please refer to this webpage for more detailed information of the topology file format.
Optionally, you can add more options to the aa_2_cg.jl command:
$ /home/user/genesis_cg_tool/src/aa_2_cg.jl 1pgb.pdb --psf --cgpdb
$ ls
1PGB.pdb 1PGB_cg.gro 1PGB_cg.itp 1PGB_cg.pdb 1PGB_cg.psf 1PGB_cg.top
The two extra files, “1PGB_cg.psf” and “1PGB_cg.pdb“, are not necessary for running MD simulations. However, you many want to use them in some structure visualization softwares. Importantly, this simplified “psf” file does not have enough information about the interaction terms, please always consider using the grotop and groitp files for simulations and analysis.
Now, let’s check the CG pdb file with VMD:
$ vmd 1PGB_cg.pdb
Change the representation to “VDW” to view the CG particles as spheres. In the following figure, we set the color of each particle according to its index. To see the same coloring effect, please change the “Coloring Method” to “Index” in VMD.
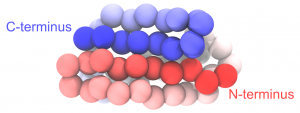
Coarse-grained protein generated from PDB 1PGB.
2. MD simulations
Now move to the simulation working directory:
$ cd 02_simulation
$ ls
param/ pro.inp
In the “param/” directory there are standard parameter files (you can find a copy from the GENESIS-CG-tool). The file “pro.inp” is the control file that contains the information for Genesis to perform the MD simulation. Let’s take a look at it:
[INPUT]
grotopfile = 1PGB_cg.top # topology file
grocrdfile = 1PGB_cg.gro # coordinate file
[OUTPUT]
pdbfile = pro_md1.pdb # PDB output
dcdfile = pro_md1.dcd # DCD trajectory
rstfile = pro_md1.rst # restart file
[ENERGY]
forcefield = RESIDCG # Residue-level CG models
electrostatic = CUTOFF # Debye-Huckel model
cg_pairlistdist_exv = 15.0 # Neighbor-list distance
[DYNAMICS]
integrator = VVER_CG # velocity-verlet propagation
nsteps = 50000000 # number of MD steps
timestep = 0.010 # timestep size (ps)
eneout_period = 10000 # energy output interval
crdout_period = 10000 # trajectory output interval
rstout_period = 100000 # restart output interval
nbupdate_period = 20 # pairlist update interval
[CONSTRAINTS]
rigid_bond = NO # don't apply constraints
[ENSEMBLE]
ensemble = NVT # Canonical ensemble
tpcontrol = LANGEVIN # Langevin thermostat
temperature = 400 # simulation temperature
gamma_t = 0.01 # thermostat friction parameter
[BOUNDARY]
type = PBC # periodic boundary condition
box_size_x = 180.0 # box size in x direction
box_size_y = 180.0 # box size in y direction
box_size_z = 180.0 # box size in z direction
The control file contains several sections, such as [INPUT], [OUTPUT], and [ENERGY], where we can specify the options for the simulation.
- In the
[INPUT]section, we set the file names for the topology file (“1PGB_cg.top“) and the coordinate file (“1PGB_cg.gro“). As described above, “1PGB_cg.top” is the main topology file and it also links the information of “1PGB_cg.itp“. - In the
[OUTPUT]section, output filenames are set. Atdyn does not create any output file unless we explicitly specify their names. Particularly, the “pdb” file contains the coordinates of the last snapshot of the simulation; the “dcd” file is the MD trajectory; and the “rst” file contains the information for restarting a simulation. - In the
[ENERGY]section, we specify the parameters related to the energy and force evaluation. RESIDCG is the name for the residue-level coarse-grained models in GENESIS. Here we also set the pair-list distance for the non-native contacts (represented by the excluded volume potentialcg_pairlistdist_exv). You can consider different values to balance computational efficiency and accuracy. As for the native contacts, all the interaction energies are calculated in GENESIS, without any cutoff. - The
[DYNAMICS]section sets up the parameters for the MD engine of atdyn. Specifically for CG simulations, GENESIS provides a high-efficiency integrator, “VVER_CG“. For the AICG2+ model, time step (timestep) can be set to 10 fs. The total number of steps (nsteps) is “50000000” in this example, but you may want to change it to a smaller value if you are carrying out the simulations on your laptop. - In the
[CONSTRAINTS]section, we disable all constraints (rigid_bond=NO). - In the
[ENSEMBLE]section, the LANGEVIN thermostat is chosen for an isothermal simulation with the friction constant of 0.01 ps-1. - Finally, in the
[BOUNDARY]section, we set the boundary conditions for the system. Generally speaking, we don’t have to worry about the computational efficiency when choosing the box size, since there is no water molecules filling the space. However, please make the box length in each dimension larger than 3 times the longestpairlistdist(57Å for the electrostatic interactions by default).
Now let’s copy necessary files prepared in the previous step:
$ cp ../01_setup/1PGB_cg.top .
$ cp ../01_setup/1PGB_cg.gro .
$ cp ../01_setup/1PGB_cg.itp .
Finally, we can run GENESIS atdyn:
$ export OMP_NUM_THREADS=2
$ mpirun -np 4 /home/user/GENESIS/bin/atdyn pro.inp > pro_md1.log
The 50000000-step simulation takes about half an hour on our PC cluster. You may want to decrease “nsteps” to save some time in your test. Once the simulation finishes, we will get four new files:
pro_md1.dcd: MD trajectory filepro_md1.pdb: PDB file of the last snapshotpro_md1.rst: MD restart filepro_md1.log: MD information log
In the “pro_md1.log” file you can check the potential energy values of each interaction term, such as “NATIVE_CONTACT” (Go-like native contact interactions) and “CG_EXV” (excluded volume interactions).
3. Analysis
We now make use of the analysis tools packaged in GENESIS to find out more information from CG MD simulations. Here we will try to compute the “Q value” from the trajectory we get in step 2.
Q values are used to estimate the “nativeness” of a conformation and have many different definitions. Here we define the Q value as the fraction of formed native contacts. When the protein is in the folded state, the Q value is high and close to 1.0; where as when the protein is unfolded, the Q value is low and close to 0.0.
Note that there are two Q-value analysis tools in GENESIS. One is called qval_analysis and is used for all-atom Go-model simulation analysis. The other is called qval_residcg_analysis, and is used for residue-level CG simulation data analysis. Here we would like to use the latter one, of course.
$ cd 03_analysis
$ ls
qval_analysis.inp
Take a look at this “qval_analysis.inp“:
[INPUT]
grotopfile = ../02_simulation/1PGB_cg.topgrocrdfile = ../02_simulation/1PGB_cg.gro[OUTPUT]qntfile = pro_md1.qval # nativeness (Q-value) output[TRAJECTORY]trjfile1 = ../02_simulation/pro_md1.dcdmd_step1 = 50000000mdout_period1 = 10000trj_format = DCD # (PDB/DCD)trj_type = COOR+BOX # (COOR/COOR+BOX)[SELECTION]group1 = all[OPTION]analysis_atom = 1 # group numberlambda = 1.2 # contact forming distance
The topology (grotopfile) and coordinate (grocrdfile) files are exactly the ones we used to perform the simulation. Native contact information are read from the topology file. For a pair of two CG particles forming native contact in the native structure, if their distance in a given conformation is within “lambda” times the native value, the contact is considered as formed between them.
/home/user/GENESIS/bin/qval_residcg_analysis qval_analysis.inp
We will get a new file containing the Q values:
$ ls
pro_md1.qval
$ head pro_md1.qval
1 0.98578
2 0.98578
3 0.96682
4 0.83886
5 0.57346
6 0.80095
7 0.83886
8 0.82464
9 0.92891
10 0.92417
Now you can use some data-visualization tools such as gnuplot to produce a figure of the time series of the Q value:
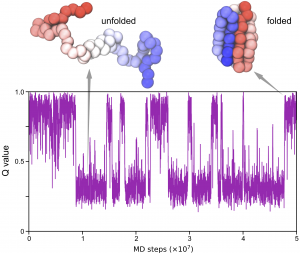
References
- Li, W., Wang, W., Takada, S., 2014, Proc. Nation. Acad. Sci., 111, 10550–10555.
-
Clementi, C., Nymeyer, H. & Onuchic, J. N., 2000, Journal of molecular biology 298, 937–953.
Written by Cheng Tan@RIKEN Center for Computational Science, Computational Biophysics Research Team
October, 2021
